Introduction
Antimicrobial resistance (AMR) develops to a state where present antibiotics can no longer effectively work against infections. AMR has become one of the most severe threats to public health because it accelerates infectious diseases that rapidly evolve into new forms.[1] Multidrug resistance (MDR) bacteria have recently added another massive burden on the healthcare system due to higher mortality associated with complications among MDR infection patients who stay long in hospitals, thus giving enough time for financial strain to set in.[2] Enterococcus faecalis, which is described as a commensal organism in the human intestine and an opportunistic pathogen of the immunocompromised [3]. The pathogenicity of E. faecalis depends on its ability to first colonize the intestinal tract and when the homeostasis is disturbed, then takes advantage of the opportunity to multiply and migrate through the lymphatics into the blood, breaching the boundary imposed on it [4]. It is still a concern for public health and has acquired multi-drug resistance, and is a source of serious problems for the treatment for hospital acquired infections. Escherichia coli (E. coli), together with enterobacter species, are among the Gram-negative bacteria of most clinical relevance in the context of a broad spectrum of hospital-acquired infections from urinary tract infections to pneumonia associated with mechanical ventilation. [5-7] Using WHO taxonomy, E. coli has been classified as a multi-resistance critical bacterial pathogen (resistant to most current marketed antimicrobial agents). [8] Escherichia coli (E. coli) is a versatile pathogen resident in the human gut; likewise, Enterococcus faecalis (E. faecalis) is recognized as a Gram-positive coccus and facultative anaerobe resident. [9,10] E. coli and E. faecalis are commensal pathogens completing the host colonization shortly after birth and thus representative of their respective phyla. [11] Methodologically, both organisms can be readily isolated, cultured, or preserved in laboratory settings. They are routinely employed as proposed indicators of human fecal contamination. Consequently, there exists an imperative to develop alternative antimicrobial agents to mitigate the issue of antibiotic resistance in bacterial populations. 1,2,3-Triazole derivatives feature a five-membered heterocyclic structure containing three nitrogen atoms and two carbon atoms. It has been a focus of considerable interest for its unique chemical and biological properties which have attracted medicinal chemists looking for effective targets or lead compounds. [12] The triazole ring is stable in various states and can adopt conformations that allow it to make diverse and synthetically accessible non-covalent interactions with biological targets [13]. Many groups can form bonding, such as dipole–dipole interactions hydrogen bonding, or π–π stacking to modulate activity of enzymes or the fit for receptor binding. [14] One of the most attractive features of the 1,2,3-triazole group is their relatively facile access during the Huisgen 1,3-dipolar cycloaddition reaction, whereby azides undergo cycloaddition with alkynes. The reaction is usually performed as a copper(I)-catalyzed “click” reaction. [15,16] This powerful transformation is characterized by its efficiency, regioselective outcome, and mild reaction conditions, upstream of a vast number of trivial variations which can lead to a great diversification in all manner of triazole derivatives. [17] An extensive literature search indicates that compounds with 1,2,3-triazole subunits have exhibited broad range of diverse activities including antiviral, anticancer, anti-inflammatory, antifungal, antibacterial. Indicating the wide applicability of the triazole as an antibiotic scaffold for multidrug resistant bacteria and for other bioactivity enhancement, the relatively easy synthetic access and high bioactivity of triazoles make them good candidates for rational drug design. [18] Specifically, wide range moieties conjugated with 1,2,3 -triazoles were reported possessing strong antibacterial properties. In this study, the problem of antibiotic resistance was addressed by designing and synthesizing new benzotriazole–triazole hybrid compounds and assessing their antibacterial activity. Bt5 and Bt6 were found as the best molecules because they showed high activities with low toxicities. Molecular docking on protein 7C7N revealed its strong binding toward important amino acids which supports its biological activity observed experimentally. The authors thus offer these compounds as potential candidates for targeting resistant bacterial strains integrating chemical synthesis, biological screening, and computation. A series of newly designed molecules were prepared and confirmed for their biological activity against selected microbial strains. Biological assessment provided an initial insight into their inhibitory potential, suggesting that both the incorporated structural features and the resulting compounds may possess valuable antimicrobial properties. Furthermore, certain computational studies were conducted to reinforce the in vitro observations and collect more knowledge on the potential interactions between the resulting agents and suitable targets. The in-silico data offered further understanding of the biology of the agents and structure-activity relationships. In conclusion, the synergetic synthetic, biological and computational undertakings accentuate the possibility of developing a drug discovery pipeline for novel antibacterial agents that can confront the so-called Antimicrobial Resistance.
Experimental Section
All reagents and solvents were commercially available (Merck, Sigma-Aldrich) and used without further purification. The reaction was monitored by thin layer chromatography using silica gel plates (60 F254); visualization was done under UV light (254 nm). Purification was carried out using column chromatography on silica gel (60 mesh) with appropriate organic solvents. Melting points were measured in a digital apparatus and are uncorrected. FT-IR spectra were recorded (Bruker spectrophotometer) using KBr discs. ¹H and ¹³C NMR spectra were recorded on a Bruker instrument (400 MHz) using DMSO- d₆ as solvent. Chemical shifts (δ) are reported in parts per million (ppm) downfield from TMS, internal standard. Mass spectra (ESI-MS) were recorded using a Thermo Scientific mass spectrometer.
Synthesis of Compound (2)
A solution of 1-(hydroxymethyl)-1H-benzo[d][1,2,3]triazole (1.0 mmol) in 15 mL dry acetone was prepared to which anhydrous K₂CO₃ (2.0-2.5 mmol) was added, and the resulting mixture was stirred at room temperature for 15 minutes. Propargyl bromide (1.2 mmol) was then added dropwise and the reaction mixture refluxed for 6 hr with constant stirring while monitoring by TLC until completion after which it is allowed to cool; filtered inorganic salts removed by filtration evaporated under reduced pressure until dryness gives a crude product. The compound (2) was obtained by column chromatography using a hexane/ethyl acetate mixture (3:1) as the eluent
1-((prop-2-yn-1-yloxy)methyl)-1H-benzo[d][1,2,3]triazole (2): pale-yellow solid, Yield: 79% ; mp 98-99 °C; IR (cm−1), 3274 (≡C–H stretch of terminal alkyne (C≡C–H), 3141(C-H of triazole ring, 3088 (aromatic C–H stretching (benzotriazole/aromatic rings), 2122 (-C≡C stretch, alkyne), 1587 (aromatic C=C); 1H NMR data δ 7.81-7.43(m, 4H, Ar-H), 5.89 (s, 2H, CH₂ attached to triazole), 4.21 (s, 2H, CH₂ linked to benzotriazole); 13C NMR data δ 144.74, 135.03, 127.63, 124.24, 119.72, 110.33, (6C, of aromatic carbons), 80.36 (1C, C≡), 75.47 (1C, ≡C–H),73.91(1C, CH₂ linked to benzotriazole), 57.59(1C, CH₂ attached to terminal alkyne).
Synthesis of 1,2,3-triazole derivatives (Bt1–Bt6)
In a round-bottom flask fitted with a magnetic stir bar, a solution containing compound 2 (1.00 mmol) and the corresponding organic azide (R–N₃, 1.10–1.50 mmol) was prepared in a mixture of ethanol and water (15 mL) at a ratio of 3:1. Following this, copper(II) sulfate pentahydrate (CuSO₄·5H₂O, 0.05 mmol, 5 mol%) and sodium ascorbate (0.20 mmol, 20 mol%) were introduced into the reaction mixture, which was then heated to 70–75 °C while stirring vigorously. The progress of the reaction was monitored using TLC (hexane/EtOAc), and after a duration of 5–6 hours, or once the alkyne had been entirely consumed, the mixture was permitted to cool to room temperature. Subsequently, the mixture was diluted with water (20 mL) and extracted with ethyl acetate (3 × 20 mL). The organic layers were combined, washed with brine, dried over anhydrous Na₂SO₄, and filtered. To eliminate any residual copper, the crude residue was either passed through a short silica gel pad or treated with a 0.05 M EDTA solution prior to the evaporation of the solvent under reduced pressure. The resulting crude product underwent purification via silica gel column chromatography, utilizing hexane/ethyl acetate as the eluent to yield the desired triazole derivatives (Bt1–Bt6) in their pure form.
1-(((1-(4-bromophenyl)-1H-1,2,3-triazol-4-yl)methoxy)methyl)-1H-benzo[d][1,2,3] triazole (Bt1): pale-yellow solid, Yield:71% , mp 187-189 °C; IR (cm−1), 3134(C-H of triazole ring), 3043 (aromatic C–H stretching, aromatic rings), 1547 (aromatic C=C); 1H NMR data δ 8.24 (s, 1H, triazole-H), 7.78-7.24 (m, 8H, Ar-H),5.85 (s, 2H, CH₂ attached to triazole), 4.70 (s, 2H, CH₂ linked to benzotriazole); 13C NMR data δ 148.14,118.89 (2C, C4 and C5 of triazole ring), 144.82, 135.11, 134.31, 133.58, 127.51, 124.32, 122.68, 121.61, 117.61, 110.14 (12C, of aromatic carbons), 73.35 (1C, CH₂ attached to triazole), 58.47 (1C, CH₂ linked to benzotriazole).
4-(4-(((1H-benzo[d](1,2,3-triazol-1-yl)methoxy)methyl)-1H-1,2,3-triazol-1-yl)benzoic acid (Bt2): White solid, Yield:74%, mp 145-147 °C; IR (cm−1), 3378 (COOH), 3154(C-H of triazole ring), 3081 (aromatic C–H stretching, aromatic rings), 1734 (carbonyl of carboxylic group), 1591(aromatic C=C); 1H NMR data δ 11.47(s, 1H, COOH), (8.28 (s, 1H, triazole-H), 7.87-7.29 (m, 8H, Ar-H),5.88 (s, 2H, CH₂ attached to triazole), 4.73 (s, 2H, CH₂ linked to benzotriazole); 13C NMR data δ 169.14 (1C, (1C, COOH), 149.49, 118.54 (2C, C4 and C5 of triazole ring), 144.81, 137.97, 135.01, 130.47, 127.49, 125.23, 124.25, 120.83, 116.93, 110.19, (12C, of aromatic carbons), 73.65 (1C, CH₂ attached to triazole), 58.97 (1C, CH₂ linked to benzotriazole).
4-(4-(((1H-benzo[d](1,2,3-triazol-1-yl)methoxy)methyl)-1H-1,2,3-triazol-1-yl)phenol (Bt3):pale-yellow solid, Yield:70%, mp 166-168 °C; IR (cm−1), 3324 (O–H stretch of phenol), 3152 (C-H of triazole ring), 3049 (aromatic C–H stretching, aromatic rings),1578(aromatic C=C); 1H NMR data δ 9.34 (s, 1H, phenol-H), 8.33 (s, 1H, triazole-H), 7.71-7.20 (m, 8H, Ar-H), 5.82 (s, 2H, CH₂ attached to triazole), 4.75 (s, 2H, CH₂ linked to benzotriazole); 13C NMR data δ 149.87,118.97 (2C, C4 and C5 of triazole ring), 157.55, 144.99, 135.02, 128.34, 127.53, 124.34, 120.67, 119.58, 117.00, 110.23, (12C, of aromatic carbons), 73.41 (1C, CH₂ attached to triazole), 58.69 (1C, CH₂ linked to benzotriazole).
4-(4-(((1H-benzo[d](1,2,3-triazol-1-yl)methoxy)methyl)-1H-1,2,3-triazol-1-yl) benzaldehyde (Bt4):White solid, Yield:73%, mp 174-176 °C; IR (cm−1), 3135 (C–H stretch of terminal aldehyde), 3166(C-H of triazole ring), 3064 (aromatic C–H stretching (aromatic rings), 1715 (carbonyl of aldehyde), 1577(aromatic C=C); 1H NMR data δ 9.89 (s, 1H, benzaldehyde-H), 8.22 (s, 1H, triazole-H), 7.79-7.25 (m, 8H, Ar-H), 5.84 (s, 2H, CH₂ attached to triazole), 4.76 (s, 2H, CH₂ linked to benzotriazole); 13C NMR data δ 187.57 (1C, benzaldehyde-C), 149.68,118.87 (2C, C4 and C5 of triazole ring), 144.94, 138.09, 135.02, 134.49, 129.92, 127.49, 124.34, 121.74, 119.44, 110.23, (12C, of aromatic carbons), 73.46 (1C, CH₂ attached to triazole), 58.98 (1C, CH₂ linked to benzotriazole).
4-(4-(((1H-benzo[d](1,2,3-triazol-1-yl)methoxy)methyl)-1H-1,2,3-triazol-1-yl)-N-(pyrimidin-2-yl)benzenesulfonamide (Bt5): Off-white solid, Yield:67%, mp 241-243 °C; IR (cm−1), 3274 (N–H stretch of sulfonamide), 3165(C-H of triazole ring), 3095 (aromatic C–H stretching, aromatic rings), 1595 (aromatic C=C); 1H NMR dat δ 11.14 (s, 1H, N-H), 8.31 (s, 1H, triazole-H), 7.97-7.04 (m, 11H, Ar-H), 5.87 (s, 2H, CH₂ attached to triazole), 4.73 (s, 2H, CH₂ linked to benzotriazole); 13C NMR data δ 143.47,118.97 (2C, C4 and C5 of triazole ring), 157.54, 156.49, 144.74, 139.24, 137.45, 135.27, 129.51, 127.47, 124.23, 121.75, 117.69, 112.76, 110.74, (15 C, of aromatic carbons), 73.46 (1C, CH₂ attached to triazole), 58.98 (1C, CH₂ linked to benzotriazole).
4-(4-(((1H-benzo[d](1,2,3-triazol-1-yl)methoxy)methyl)-1H-1,2,3-triazol-1-yl)-N-(5-methylisoxazol-3-yl)benzenesulfonamide (Bt6):pale-white solid, Yield:69 %, mp 241-243 °C; IR (cm−1), 3263 (N–H stretch of sulfonamide), 3114(C-H of triazole ring), 3072 (aromatic C–H stretching, aromatic rings), 1579 (aromatic C=C); 1H NMR data δ 11.22 (s, 1H, N-H), 8.36 (s, 1H, triazole-H), 7.91-7.18 (m, 13H, Ar-H), 6.11(s, 1H, CH, isoxazole), 5.83 (s, 2H, CH₂ attached to triazole), 4.70 (s, 2H, CH₂ linked to benzotriazole), 2.31(s, 3H, CH3 attached to isoxazole), ; 13C NMR data δ 169.24, 156.54, 96.87 (3C, C5, C3 and C4 of isoxazole ring), 142.98,118.91(2C, C4 and C5 of triazole ring), 144.87, 139.68, 137.79, 135.28, 129.65, 127.87, 124.29, 121.75, 120.55, 112.27, (10C, of aromatic carbons), 73.78 (1C, CH₂ attached to triazole), 58.61 (1C, CH₂ linked to benzotriazole), 13.14 (1C, CH3 linked to isoxazole).
Biological assays
Antibacterial evaluation of the prepared 1,2,3-triazole compounds (Bt1-Bt6) was carried out in vitro against selected four bacterial strains Enterococcus faecalis, clinical Enterococcus faecalis isolate, Escherichia coli, and clinical Escherichia coli isolate. All tested strains were sourced from (ATCC). Fresh susceptibility test organisms were prepared in normal saline just prior to screening. The minimum inhibitory concentration (MIC) of each compound was determined employing the microdilution method. Compounds were serial ten-fold diluted in microtiter plates, and incubated overnight at 37 °C after the addition of a standardized bacterial inoculum (1-2 × 10⁷ CFU/mL; 10 μL per well). The MIC was based on the lowest concentration at which there was no visible bacterial growth (an absence of turbidity compared to control wells) using ciprofloxacin as drug of reference. Thus, the results detailed the antibacterial activity of the novel benzotriazole–triazole derivatives against the selected strains.
Results Discussion
Compound (2), 1-((prop-2-yn-1-yloxy)methyl)-1H-benzo[d][1,2,3]triazole was synthesized via a nucleophilic substitution reaction between 1-(hydroxymethyl)-1H-benzo[d][1,2,3]triazole and reacted with propargyl bromide using anhydrous potassium carbonate. Dry acetone and mild basic conditions were chosen for the easy deprotonation of the hydroxyl group to form alkoxide which would undergo an SN2 reaction with propargyl bromide to give the desired propargyl ether (Fig. 1). The IR spectrum showed that the etherification was successful together with alkyne moiety present as shown by sharp bands at 3274 cm⁻¹(≡C–H stretching)and 2122 cm⁻¹(C≡C stretch). Aromatic, triazole C–H stretching bands appeared between 3088–3141 cm⁻¹. The formation of compound (2) is further evidenced by its ^1H NMR spectrum showing singlet peaks due to two methylene groups attached respectively on triazole ring and benzotriazole core observed at δ values5 .89and4 .21ppm, The ¹³C NMR spectrum appears in good agreement with the proposed structure, especially signals at δ 80.36 and 75.47 ppm which are characteristic of terminal alkyne carbons.[16] The synthesized alkyne intermediate (2) was a key precursor for Cu(I)-catalyzed azide–alkyne cycloaddition (CuAAC) reaction to give a series of 1,4-disubstituted triazole derivatives (Bt1–Bt6). Click reaction in ethanol/water medium under CuSO₄·5H₂O/sodium ascorbate catalytic system efficiently generates Cu(I) species in situ allowed the regioselective [17,18] formation of 1,4-substituted 1,2,3-triazoles. TLC monitoring revealed very efficient cycloaddition giving yields in the range 67–74% with different azide substrates showing suitability of reaction conditions toward a wide range of azide substrates.
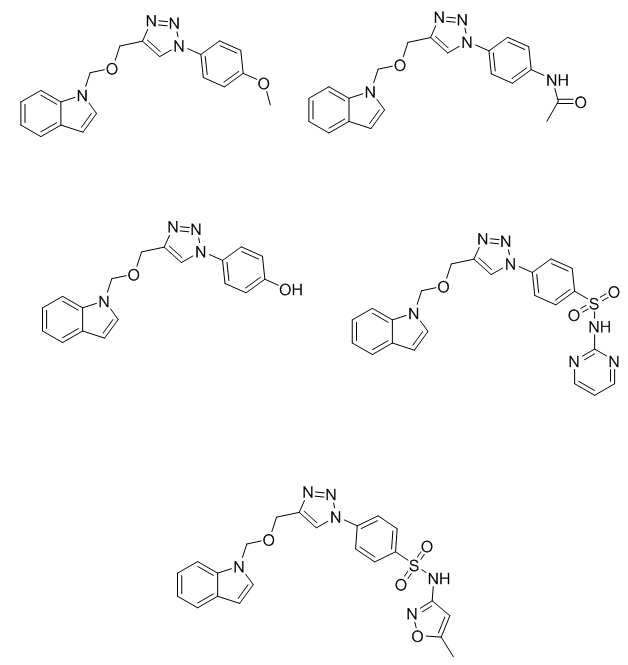
Fig.1: Synthesis of 1,2,3- triazole derivatives (Bt1-Bt6)
The spectral data of the derivatives confirmed the cycloaddition. A triazole C-H singlet was observed at δ 8.2-8.3 ppm in the ¹H NMR spectra of all the compounds along with aromatic multiplets due to benzotriazole and aryl/heteroaryl substituents. The methylene bridges between benzo-triazole and triazole rings appeared as singlets at about δ 5.82-5.88 ppm and 4.70-4.76 ppm, respectively. In their IR spectra, disappearance of alkyne stretches (3274 and 2122 cm⁻¹) together with presence both triazole as well aromatic C-H bands proved full cycloaddition. Each derivative showed also diagnostic functional-group absorptions coherent its structure: COOH stretch Bt2 (3378&1734), phenolic O–H band Bt3(3324), aldehydic C=O Bt4(1715). For sulfonamide containing derivatives (Bt5&Bt6) strong N–H stretches around3263–3274 together additional aromatic signals confirm successful incorporation sulfonamide moiety. The NMR spectra further revealed differences in the structures of the various derivatives. For example, Bt6 displayed extra signals at δ 6.11 and 2.31 ppm attributable respectively to the isoxazole proton and its methyl substituent consistent with heterocyclic substitution. In all cases, the ¹³C NMR spectra showed triazole carbons around δ 118–149 ppm together with aromatic carbon distributions agreeing with proposed structures. Unique carbonyl, heteroaryl or substituent specific signals provided additional evidence for each compound’s identity. IR and NMR data clearly indicated successful synthesis not only of propargyl benzotriazole intermediate but also its subsequent conversion into a diverse library of 1,4-disubstituted 1,2,3-triazole derivatives via CuAAC as reflected by high yields accompanied by clean spectral profiles plus stability observed for obtained compounds thereby demonstrating this synthetic approach to be highly efficient besides being versatile in generating multifunctional benzotriazole–triazole hybrid molecules.
Antibacterial Activity
The antibacterial activities of the benzotriazole–triazole hybrids (Bt1–Bt6) were evaluated against selected strains of bacteria in relation to their potency. It was found that the activities of the compounds were dependent on the nature of the substituents on the triazole, and moreover their electronic nature leading to a change in the molecular structure of the benzotriazole and triazole, as summarized in Table 1. Most of their derivatives are broad spectrum antibacterial agents, revealing that benzotriazole–triazole hybrid scaffold represent a valid ‘space’ or scaffold for antimicrobial drug design. The rest of their agents, even had comparable and lower MICs to Ciprofloxacin in selected strains. Moreover, initial structure–activity relationship (SAR) investigations indicated that substituents that are electron withdrawing or donating on the triazole ring would influence antibacterial activity by modulating electronic distribution, lipophilicity and capacity to bind to enzymes or cellular components in bacteria. Taken together, these results suggest that benzotriazole-triazole hybrids are a promising new class of antibacterial agents. Overall, the most active analogues identified in the course of this study represent good starting points in the search for drug leads against drug-resistant infections.
Table 1. Antibacterial efficacy (MIC, µg/mL) and cytotoxic profile (mg/mL) of prepared compounds Bt1–Bt6
| Com. no. | Gram + | Gram − | CC50 value (µg mL−1) | ||
| E. faecalis | E. faecalis (Clinical isolate) | E. coli | E. coli (Clinical isolate) | ||
| Bt1 | 12.25 | 51 | 18.24 | 5.87 | 45 |
| Bt2 | 18.57 | 50 | 31.24 | 15.58 | 70 |
| Bt3 | 8.98 | 28.36 | 1.85 | 8.57 | 95 |
| Bt4 | 3.14 | 5.57 | 2.35 | 3.24 | 60 |
| Bt5 | 0.49 | 7.24 | 0.24 | 1.04 | 120 |
| Bt6 | 0.57 | 11.20 | 0.47 | 1.24 | 135 |
| Ciprofloxacin | 0.78 | 12.5 | 0.12 | 0.78 | – |
| Chloroquine | – | – | – | – | >100 |
Bt5 and Bt6 demonstrated the most significant antibacterial efficacy within the series, particularly against the Gram-negative bacterium (E. coli and a clinical isolate). Bt5 exhibited remarkable potency with (MIC) values of 0.24 µg/mL and 1.04 µg/mL, whereas Bt6 recorded comparably low MIC values of 0.47 µg/mL and 1.24 µg/mL, respectively. These are very close to ciprofloxacin (0.12–0.78 µg/mL) thus indicating that the introduction of sulfonamide heterocycles (pyrimidine in Bt5 and isoxazole in Bt6) has a major effect on antimicrobial potency.[19] Their activities against Gram-positive E. faecalis strains followed the same trend with Bt5 (0.49 µg /m L ) being the most active derivative.[20] The results show that sulfonamide functionality leads to better interaction with bacterial enzymatic targets presumably by way of hydrogen bonding or due to increased polarity. Bt4 showed moderate activity with MIC values ranging from 2.35 to 5.57 µg/mL, thus emphasizing that the presence of an aldehyde group increases its electron-withdrawing character and makes the antibacterial action better than Bt1-Bt3. Compounds Bt1, Bt2, and Bt3 manifested much weaker antimicrobial effects, particularly against the E. faecalis clinical isolate wherein MIC values were recorded at 28-50 µg/mL; this clearly indicates that very simple phenyl or carboxylic acid or phenolic substituents provide insufficient enhancement to antibacterial activity.
Cytotoxicity Studies CC₅₀ values indicated that most of the compounds were not significantly toxic, mainly Bt5 (120 µg/mL) and Bt6 (135 µg/mL), with maximum CC₅₀ values recorded for the series (Fig. 2). This result is quite indicative of a wide safety margin and good selectivity toward bacterial cells relative to mammalian cells. Bt3 also exhibited a high CC₅₀ (95 µg/mL); Bt1, Bt2, and Bt4 displayed moderate cytotoxicity (45-70 µg/mL). Interestingly, the two most active antibacterial compounds were also found to be least toxic thus exhibiting an excellent therapeutic index. On comparing synthesized derivatives with reference antibiotics; ciprofloxacin still remained more potent but only slightly so when compared with either Bt5 or Bt6.Chloroquine showed very weak antibacterial activity hence was used primarily as a reference for cytotoxicity. A clear SAR tendency is revealed by the combined results: heterocyclic sulfonamide substituents markedly enhance antibacterial activity while maintaining low toxicity, whereas simpler electron-donating or neutral substituents result in weaker activity. Bt5 and Bt6 can be considered the best antibacterial candidates of this series because they are not only highly effective-and their effects cover a broad spectrum-but also safe to a high degree. Further development and fine-tuning studies may thus follow based on these two compounds.
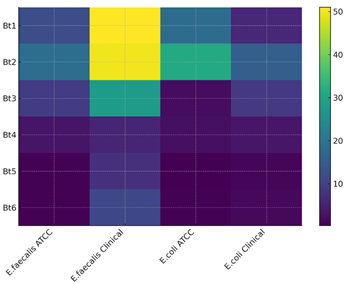
Fig. 2: Heatmap of Bt1–Bt6 Bioactivity Against Clinical and ATCC Strains of E. faecalis and E. coli.
Validation of Docking Protocol
Figure 3 indicates the redocking of the native co-crystallized ligand into the protein’s active site 7C7N to validate the docking protocol. An RMSD of 0.1324 Å was obtained for the redocking, indicating an excellent fit between predicted docking pose and the crystallographic conformation. The very small deviation between crystallographic pose (green) and redocked pose (red) indicates good correspondence of docking protocol and crystallographic binding mode. Importantly, the redocked ligand retained key interactions with major active site residues, including Asp49, Asp73, Pro79, Asn46, Gly77, Arg76, and Arg136, through a combination of hydrogen bonding and electrostatic contacts. The preservation of these conserved interactions suggests that the docking parameters and scoring function employed in this study realistically represent the physicochemical environment of the active site of protein 7C7N. The capacity of this protocol to reproduce binding orientation and key interactions, significantly supports the reliability and robustness of our setup. Hence low RMSD and conservation of interactions in some critical residues stimulate confidence in our method for the docking studies of designed benzotriazole–triazole derivatives.
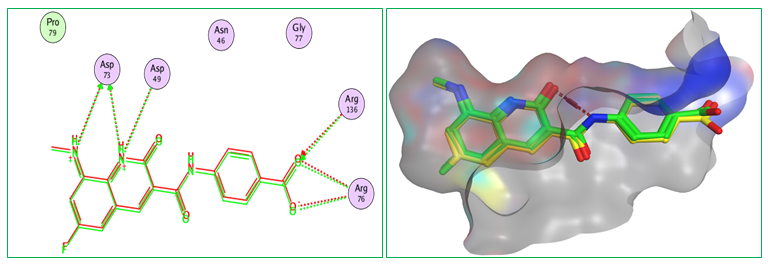
Fig. 3: Validation of the Docking Protocol for Protein 7C7N Using the Native Ligand (RMSD = 0.1324 Å
Molecular Docking Studies
Binding affinities and interactions of Bt1–Bt6 ligands show different behaviors because their structures are somewhat different, as seen from results scores on the 7C7N protein. Bt5 recorded the best or lowest docking score among all ligands at –7.4426 kcal·mol⁻¹; hence, it can be said to have the highest binding affinity with a high number of hydrogen bond donor-acceptor interactions involving Arg76 together with π–H contacts involving Ile78 and Pro79 that stabilize its orientation within the binding pocket. The other two compounds which also gave good docking scores were Bt2 (–7.3436) & Bt1(–7 .1984); both formed important stabilizing hydrogen bonds Asp73/Val71(involved in π–H interaction)/Ile78. Bt3, Bt4, and Bt6 recorded slightly weaker binding energies relative to the best ligands but they made relevant interactions with important residues in the active site. Bt3 and Bt4 made more contacts with Asp49 and Asp73 which means that acidic residues are very instrumental in holding these ligands as shown in Figures 6 and 7. With a moderate docking score of -6.8270 kcal·mol⁻¹, Bt6 formed the most varied interaction profile comprising hydrogen bond acceptor and donor interactions with Arg76, Arg136, Met95 together with a strong π–cation interaction with Arg76 (Figure 9).
Table 1. Molecular docking scores and binding interactions of ligands (Bt1-Bt6) against (PDB: 7C7N)
| Ligand | RMSD (Å) | Dock Score (E, kcal·mol⁻¹) | Interaction type | ΔE (kcal·mol⁻¹) per interaction | Distance (Å) | Residue / Partner |
| Bt1 | 1.9964 | -7.1984 | H-B-D | -1.7 | 3.40 | VAL 71 |
| pi-H | -0.7 | 3.70 | ILE 78 | |||
| pi-H | -0.6 | 4.52 | ILE 78 | |||
| Bt2 | 0.9776 | -7.3436 | H-B-D | -1.0 | 3.25 | ASP 73 |
| H-B-D | -0.7 | 3.20 | VAL 71 | |||
| Bt3 | 1.4186 | -6.8305 | H-B-D | 3.26 | -0.8 | ASP 49 |
| Bt4 | 1.2893 | -6.9847 | H-B-D | 3.18 | -0.7 | ASP 73 |
| pi-H | 4.28 | -0.9 | ILE 78 | |||
| pi-H | 3.68 | -1.5 | PRO 79 | |||
| Bt5 | 1.9269 | -7.4426 | H-B-D | 2.96 | -2.4 | ARG 76 |
| H-B-A | 3.09 | -2.9 | ARG 76 | |||
| pi-H | 4.20 | -1.4 | ILE 78 | |||
| pi-H | 3.58 | -0.7 | PRO 79 | |||
| Bt6 | 1.9124 | -6.8270 | H-B-D | 4.07 | -0.3 | MET 95 |
| H-B-A | 3.31 | -1.4 | ARG 136 | |||
| H-B-A | 3.34 | -0.9 | ARG 136 | |||
| H-B-A | 3.01 | -1.1 | ARG 76 | |||
| pi-cation | 3.63 | -1.8 | ARG 76 | |||
| pi-H | 3.78 | -1.3 | ILE 78 |
* Note: hydrogen bond donor=H-B-D; hydrogen bond acceptor=H-B-A
The interactions show a wider binding footprint that could be responsible for its moderate yet stable docking score. The key binding residues highlighted in the ligand interaction profiles are Arg76, Ile78, Pro79 and also Asp73 plus Val71. Hydrogen bonds as well as hydrophobic or π interactions seem important for strong binding of a ligand to this protein wherein Bt5 is indicated by its network synergy effect among interactions formed within it into being most likely candidate binder towards 7C7N protein target used here. Ligands occupy about same region inside active site but oriented differently according structural features (see two/ three-dimensional interaction diagrams; Figures: 4-9) thus Bt-series ligands could bind effectively against seven proteins meanwhile guiding future optimization based on structure towards drug design.
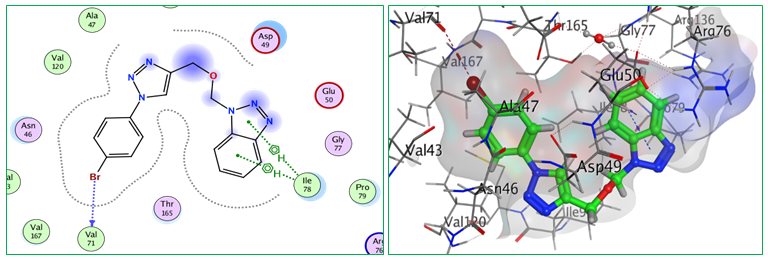
Fig.4: 2D and 3D representations of ligand interactions (Bt1) with active sit of (7C7N).
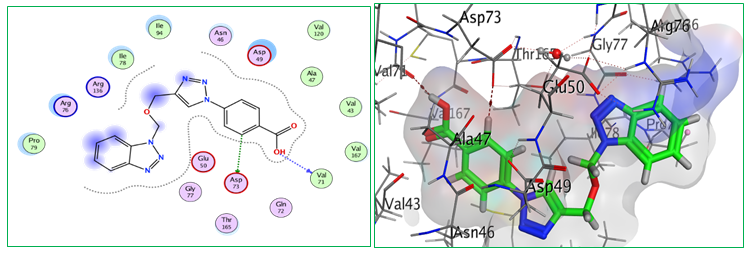
Fig.5: 2D and 3D representations of ligand interactions (Bt2) with active sit of (7C7N
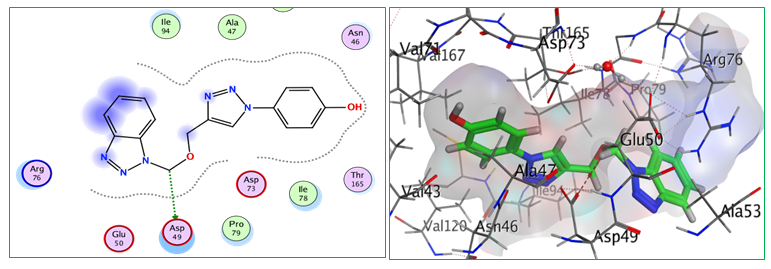
Fig.6: 2D and 3D representations of ligand interactions (Bt3) with active sit of (7C7N).
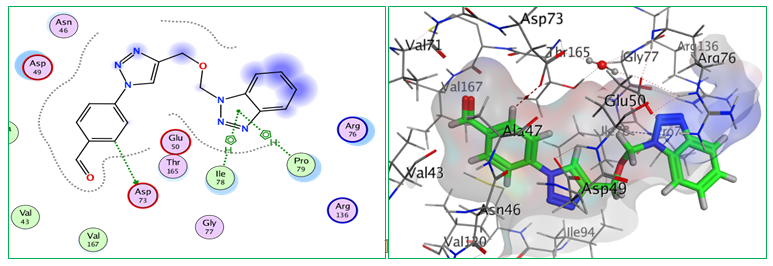
Fig.7: 2D and 3D representations of ligand interactions (Bt4) with active sit of (7C7N).
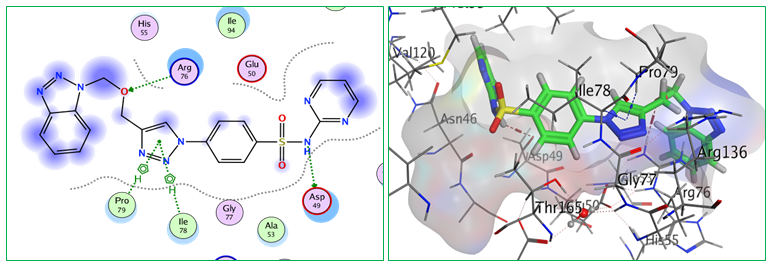
Fig.8: 2D and 3D representations of ligand interactions (Bt5) with active sit of (7C7N)
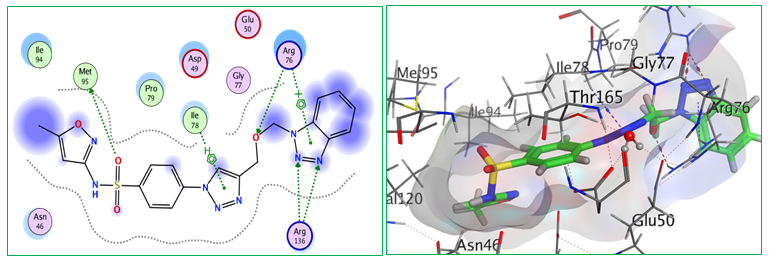
Fig.9: 2D and 3D representations of ligand interactions (Bt6) with active sit of (7C7N).
Conclusion
A novel series of benzotriazole–1,2,3-triazole hybrid derivatives (Bt1–Bt6) was synthesized by Cu(I)-catalyzed azide-alkyne cycloaddition reaction. The structures were elucidated using different spectroscopic techniques. Antibacterial results showed that molecular modifications at the triazole moiety assumed an important role in its biological activity; Bt5 and Bt6 exhibited minimum and broadest antibacterial activities among the synthesized compounds with MIC values against most bacteria comparable to ciprofloxacin, a standard drug, besides excellent selectivity manifested through low cytotoxicity hence indicating high antimicrobial effect brought about by sulfonamide linked heterocycles. Molecular docking study supported antibacterial experimental results since two compounds Bt5 & Bt6 had good binding scores against 7C7N protein target where compound Bt5 was found having best docking score forming many stable interactions with important active site residues. The docking protocol was validated by redocking of the native ligand, giving a very low RMSD value (0.1324 Å). This supports the reliability of this computational approach. In summary, benzotriazole–triazole hybrid compounds Bt5 and Bt6 can be considered promising lead scaffolds for developing new antibacterial agents to act against multidrug-resistant pathogens discovered through studies reported herein. Further optimization on these compounds as well as detailed mechanism study is warranted before pushing them towards preclinical evaluation.
References
- Dadgostar, Pari. “Antimicrobial resistance: Implications and costs.” Infection and Drug Resistance 12 (2019): 3903–3910.
- Llor, Carl, and Lars Bjerrum. “Antimicrobial resistance: risk associated with antibiotic overuse and initiatives to reduce the problem.” Therapeutic Advances in Drug Safety 5.6 (2014): 229–241.
- Rigottier-Gois, L., Madec, C., Navickas, A., Matos, R. C., Akary-Lepage, E., Mistou, M. Y., & Serror, P. “The surface rhamnopoly saccharide Epa of Enterococcus faecalis is a key determinant of intestinal colonization.” Journal of Infectious Diseases 211.1 (2015): 62–71.
- Manoharadas, S., Altaf, M., Ahmad, N., Alrefaei, A. F., & Al-Rayes, B. F. “Construction and activity testing of a modular fusion peptide against Enterococcus faecalis.” Antibiotics 12.388 (2023): 1–12.
- Zhou, X. M., Hu, Y. Y., Fang, B., & Zhou, C. H. “Benzene sulfonyl thiazolo imines as unique multitargeting antibacterial agents towards Enterococcus faecalis.” European Journal of Medicinal Chemistry 248 (2023): 115088.
- Muller, C., Massier, S., Le Breton, Y., & Rincé, A. “The role of the CroR response regulator in resistance of Enterococcus faecalis to D-cycloserine is defined using an inducible receiver domain.” Molecular Microbiology 107.3 (2018): 416–427.
- Ruppé, E., Woerther, P.-L., & Barbier, F. “Mechanisms of antimicrobial resistance in Gram-negative bacilli.” Annals of Intensive Care 5 (2015): 21–36.
- World Health Organization. Prioritization of pathogens to guide discovery, research and development of new antibiotics for drug-resistant bacterial infections, including tuberculosis. WHO Press, Geneva, 2017.
- Mohamed, J. A., & Huang, D. B. “Biofilm formation by enterococci.” Journal of Medical Microbiology 56.11 (2007): 1581–1588.
- Tenaillon, O., Skurnik, D., Picard, B., & Denamur, E. “The population genetics of commensal Escherichia coli.” Nature Reviews Microbiology 8.3 (2010): 207–217.
- Jimenez, E., Marin, M. L., Martin, R., Odriozola, J. M., Olivares, M., Xaus, J., Fernandez, L., & Rodriguez, J. M. “Is meconium from healthy newborns actually sterile?” Research in Microbiology 159.3 (2008): 187–193.
- Rostovtsev, V. V., Green, L. G., Fokin, V. V., & Sharpless, K. B. “A stepwise Huisgen cycloaddition process: Copper(I)-catalyzed regioselective ligation of azides and terminal alkynes.” Angewandte Chemie International Edition 41.14 (2002): 2596–2599.
- Kolb, H. C., & Sharpless, K. B. “The growing impact of click chemistry on drug discovery.” Drug Discovery Today 8.24 (2003): 1128–1137.
- Agalave, S. G., Maujan, S. R., & Pore, V. S. “Click chemistry: 1,2,3-triazoles as pharmacophores.” Chemistry – An Asian Journal 6.10 (2011): 2696–2718.
- Huisgen, R., Szeimies, G., & Möbius, L. “1,3‐Dipolar cycloadditions, XXXII. Kinetics of the addition of organic azides to C≡C multiple bonds.” Chemische Berichte 100.8 (1967): 2494–2507.
- Massarotti, A., Aprile, S., Mercalli, V., Di Grosso, E., Grosa, G., Sorba, G., & Tron, G. C. “Are 1,4- and 1,5-disubstituted 1,2,3-triazoles good pharmacophoric groups?” ChemMedChem 9.12 (2014): 2497–2508.
- Agalave, S. G., Maujan, S. R., & Pore, V. S. “Click chemistry: 1,2,3-triazoles as pharmacophores.” Chemistry – An Asian Journal 6.10 (2011): 2696–2718.
- Gonzalez-Calderon, D., Mejía-Dionicio, M. G., Morales Reza, M. A., Ramírez-Villalva, R., Morales-Rodríguez, M., Jauregui-Rodríguez, B., Díaz-Torres, E., Gonzalez-Romero, C., & Fuentes-Benítes, A. “Azide–enolate 1,3-dipolar cycloaddition in the synthesis of novel triazole-based miconazole analogues as promising antifungal agents.” European Journal of Medicinal Chemistry 112 (2016): 60–65.
- Kant, R., Singh, V., Nath, G., Awasthi, S. K., & Agarwal, A. “Design, synthesis and biological evaluation of ciprofloxacin-tethered bis-1,2,3-triazole conjugates as potent antibacterial agents.” European Journal of Medicinal Chemistry 124 (2016): 218–228.
- Singh, M. K., Tilak, R., Nath, G., Awasthi, S. K., & Agarwal, A. “Design, synthesis and antimicrobial activity of novel benzothiazole analogs.” European Journal of Medicinal Chemistry 63 (2013): 635–644.
- Agarwal, A., Singh, P., Maurya, A., Patel, U. K., Singh, A., & Nath, G. “Ciprofloxacin-tethered 1,2,3-triazole conjugates: New quinolone family compounds to upgrade our antiquated approach against bacterial infections.” ACS Omega 7 (2022): 2725–2736.
